Once again, one of very old Katka’s dataset was published. This time, the blood samples which she collected from Sericornis birds from along elevational gradient of Mt. Wilhelm were used in collaboration with Frank Rheindt.
In our recent work, we analyzed thousands of genome-wide DNA markers across an assemblage of three closely related understorey-inhabiting scrubwrens (Sericornis and Aethomyias; Aves) from montane forest along an elevational gradient on Mt. Wilhelm, the highest mountain of Papua New Guinea. Despite species-specific differences in elevational preference, we found limited differentiation within each scrubwren species, but detected a strong genomic signature of simultaneous population expansions at 27-29 ka, coinciding with the onset of the LGM.
The remarkable synchronous timing of population expansions of all three species demonstrates the importance of global cooling cycles in expanding highland habitat. Global cooling cycles have likely had strongly different impacts on tropical montane areas versus boreal and temperate latitudes, leading to population expansions in the former and serious fragmentation in the latter.
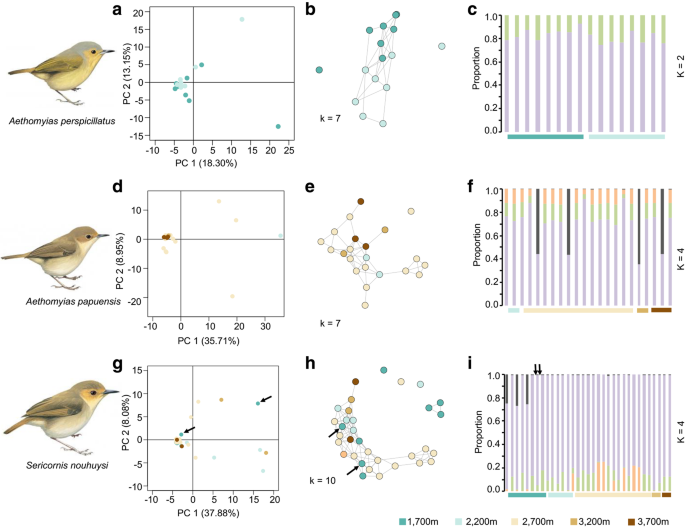
Population-genomic subdivision in the three species of scrubwren based on principal coordinate analysis (left), network-based analysis (middle) and STRUCTURE analysis (right). See Table 1 for the number of SNPs used in each analysis. The two S. nouhuysi individuals from the Finisterre Range are marked with arrows. Bird illustrations modified from del Hoyo et al. [23]
Ehm, one of the “specimen” in Katka’s dirty hand (Sericornis nouhuysi from Bananumbu cam)